Traces the origin of an antibiotic resistance gene in the hope of blocking its transmission
“Superbug” resistant to strong antibiotics is a ticking time bomb in global public health. A joint study led by a scientist from City University of Hong Kong (CityU) has traced the potential origin of an antibiotic resistance gene and identified a type of bacteria as the possible source. The team also studied the gene’s transmission mechanisms and published their findings recently, in the hope of blocking its transmission.
The research team was led by Professor Chen Sheng, Associate Dean (Research) of Jockey Club College of Veterinary Medicine and Life Sciences (JCC) as well as Professor of Veterinary Bacteriology at CityU, together with experts from mainland China. Their findings were published in the scientific journal Nature Communications, titled “Epidemiological and phylogenetic analysis reveals Flavobacteriaceae as potential ancestral source of tigecycline resistance gene tet(X)”.
Antibiotic resistance: a global threat
Antibiotic resistance has already been listed by the World Health Organization (WHO) as “one of the biggest threats to global health, food security, and development today”. Antibiotics are used to treat bacterial infections. However, the overuse of them has led to the evolution of bacteria that exhibit antibiotic resistance. As a result, related infections have become difficult to be treated and can eventually prolong the hospitalisation period and increase the death rate.
Even worse, the Covid-19 pandemic has escalated the issue. Dr Tedros Adhanom Ghebreyesus, Director-General of WHO, pointed out in June 2020 that the Covid-19 pandemic has led to increased use of antibiotics, which will lead to higher bacterial resistance rates.
Professor Chen explained one of the reasons: frontline doctors would prescribe antibiotics to Covid-19 patients to prevent other infections, therefore the use of antibiotics would increase. He pointed out that doctors’ prescriptions usually start with the most commonly used first-line antibiotics. If the treatment failed, then doctors would use the stronger second-line drugs. However, in many places around the world, first-line antibiotics have already failed to treat infections, even second or third-line antibiotics are not effective anymore. Doctors are facing a difficult situation of having no effective antibiotics to prescribe.

Professor Chen also pointed out that the problem of antibiotic resistance in mainland China is serious. Medical staffs might need to prescribe different antibiotics in one go, for example using tigecycline and carbapenem together to treat patients. “Usually carbapenem does not have effects on patients in China anymore. Therefore tigecycline is considered as the last resort to treat serious infections caused by multi-drug resistant bacteria,” he said.
Yet the increased usage of tigecycline has led to the problem of resistance, too. There have been sporadic tigecycline-resistance cases reported worldwide, including in mainland China. After study, scientists have discovered a group of tigecycline-resistance tet(X) gene. Tigecycline would be much less effective on patients who are infected with bacteria carrying such a gene. However, scientists still do not know much about tet(X) gene.
Potential origin of tigecycline-resistance gene
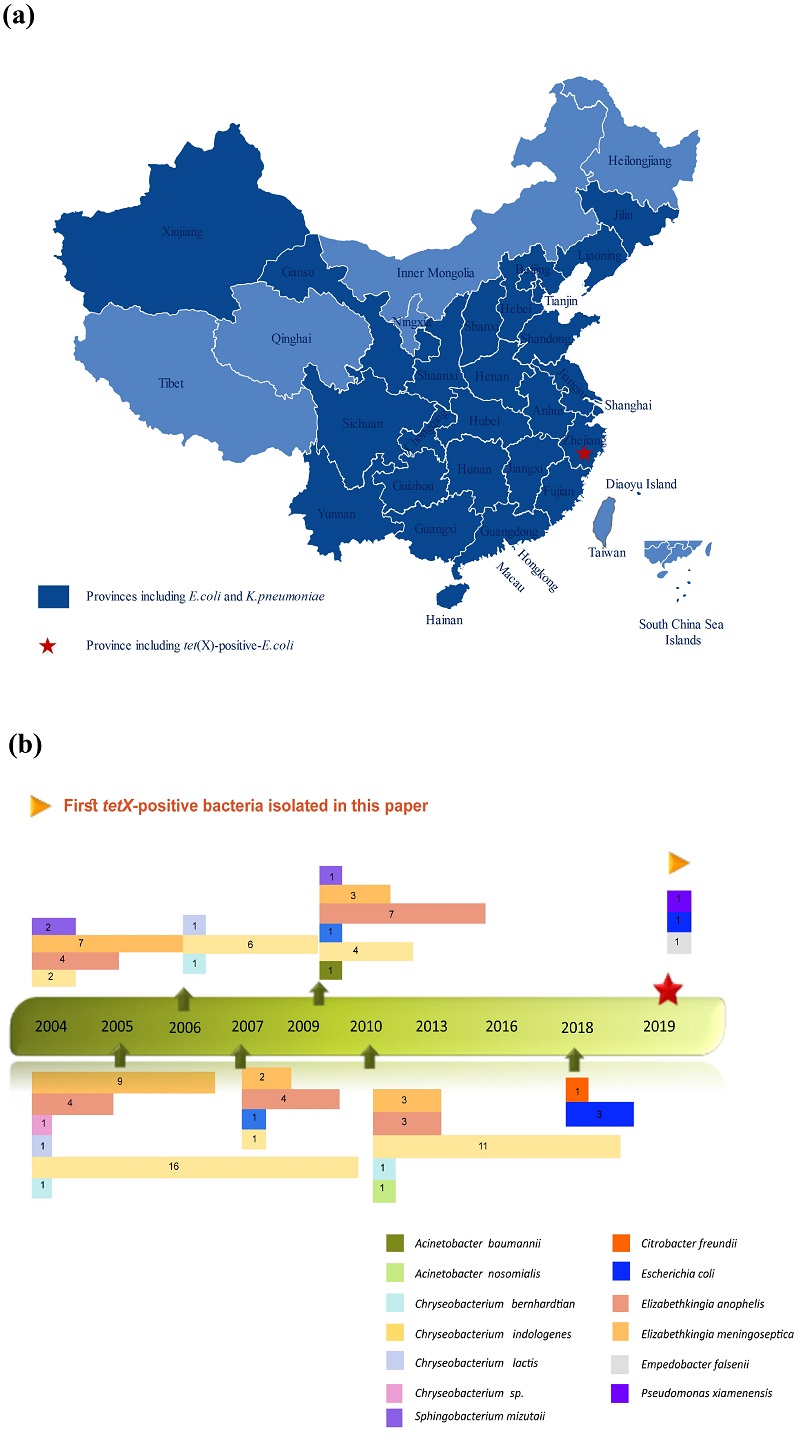
Professor Chen’s team researched on tet(X) and discovered for the first time, its potential origin and transmission mechanisms. They collected 2,997 clinical samples of E. coli and K. pneumoniae from 77 hospitals in 26 provinces in China. The tet(X) and its orthologues were found in 4 (0.32%) E. coli samples and none were found in the K. pneumoniae clinical samples.
Since the four tet(X)-bearing E. coli samples were all collected from the Zhejiang province, the team carried out further surveillance of tet(X) orthologues in 3,692 clinical Gram-negative bacterial strains collected in hospitals in Zhejiang province from 1994 to 2019. A total of 102 tet(X)-bearing strains were found, and 95 of them were Flavobacteriaceae, the dominant bacteria. All the 95 strains showed resistance to multiple antibiotics, including tigecycline.
The whole-genome sequencing and bioinformatics analysis were performed on these strains, and the team discovered that tet(X) were mainly located on the chromosomes of Flavobacteriaceae, but not the plasmid. “This indicated that the gene is probably present in Flavobacteriaceae naturally. In fact, many of these resistance genes exist in the natural ecosystem. It is just a coincidence that tigecycline is used and tet(X) so happened to play a role, therefore bacteria with this gene could survive,” explained Professor Chen.
The team also discovered that tet(X) shared similar GC-content of about 37% as Flavobacteriaceae. It means both genes were very similar. GC-content refers to the percentage of nucleotides in the genomic sequence that are either guanines (G) or cytosines (C), two chemical components of DNA. Organisms are found to have specific GC-content which could be calculated from a fragment of DNA, RNA or entire genome. According to the above findings from this research, the team suggested that Flavobacteriaceae could be the potential ancestral source of the tigecycline resistance gene tet(X).
The study also showed that the original tet(X) genes found on the chromosomes of Flavobacteriaceae have further evolved. “According to the genome analysis data, we found DNA fragment of tet(X) in Enterobacteriaceae, Acinetobacter as well as Flavobacteriaceae, which indirectly indicated that genetic exchange could occur among different species,” elaborated Professor Chen.
Blocking the transmission of tet(X)
“From this study, we have discovered the potential source of tet(X) and how the gene would evolve. These important information could control its spread clinically. In the future, we should also pay more attention to the ‘source’ Flavobacterium and whether tet(X) has become more prevalent. If so, that means the mechanism of gene transmission may have changed,” concluded Professor Chen.
His team will conduct follow-up researches, and team members are currently working on the systematic classification of the variants of tet(X) genes according to their functions and characteristics, which would facilitate future related researches.

Professor Chen and Professor Chen Gongxiang from the Second Affiliated Hospital of Zhejiang University, are the corresponding authors of the paper. The co-first authors are Dr Dong Ning, Research Assistant of JCC at CityU, Professor Zhang Rong from the Second Affiliated Hospital of Zhejiang University, and Professor Shen Zhangqi of China Agricultural University. Research team members from CityU also included Research Assistant Cheng Qipeng. Other researcher comes from The Hong Kong Polytechnic University.
The study received funding support from the National Natural Science Foundation of China.
DOI number: 10.1038/s41467-020-18475-9
Related story: A weapon to make a superbug to become more deadly