- 1A-303, 3/F, Block 1, To Yuen Building
- +852 3442-2914
- +852 3442-0549
- peter.lidsky@cityu.edu.hk
- CityUHK Scholars
- Lab Website
- aging • evolution • virology • cellular senescence • Drosophila
Prof. Peter Lidsky received his MS and PhD training in virology at Moscow State University, where his research focused on virus-host interactions in Picornaviruses. He then transitioned to the University of Zurich to develop tools and probes for fluorescent imaging in the Drosophila model. Finally, at the University of California San Francisco (UCSF), he combined his expertise in virology and genetics to build a versatile research program. Notably, at UCSF, he developed the “pathogen control” hypothesis of aging, which forms the foundation of his current research interests.
Fighting Aging: Building Upon Evolutionary Theory
Aging is an evolutionary paradox: it remains unclear why it evolved, and there is still no consensus on its fundamental nature. The debate over whether aging is a programmed genetic adaptation or a passive consequence of accumulated damage has persisted for more than a century. For decades, programmed aging theories were marginalized as incompatible with mainstream evolutionary thinking, while damage-centered theories dominated the field.
Recently, Prof. Lidsky proposed the pathogen control hypothesis of aging, which situates aging as an adaptive, programmed trait within standard evolutionary theory. This framework has helped re-establish programmed aging as a serious contender in explaining the biology of aging. Our lab is working to further develop this model and use it to investigate core mechanisms linking aging, immunity, and infection.
In particular, we focus on the antiviral functions of cellular senescence. While it has long been recognized that viruses can induce cellular senescence and that senescent cells can exert antiviral effects, the underlying mechanisms remain poorly characterized. We are testing the “immune militia” hypothesis of cellular senescence, in which senescent cells function as “militiamen” temporarily recruited from their normal roles to defend the organism during infection. In this model, senescent cells become resistant to viral replication, secrete factors that enhance antiviral defenses in neighboring cells, and coordinate with immune cells—the “professional soldiers” of the immune system.
Because senescent cells can be harmful if not efficiently cleared, we are also investigating whether virus-induced cellular senescence contributes to the long-term complications of viral infections, including chronic inflammation, tissue dysfunction, and accelerated aging phenotypes.
Lectures and Seminars
Interviews
Book – “Aging: Why Does Evolution Kill?”
Partner: Open Longevity
SINGULARIS: changing the ways we interact with knowledge
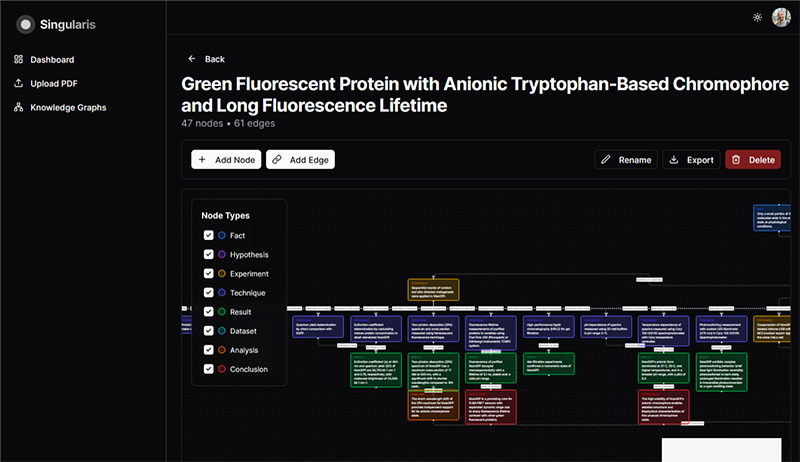
We argue that traditional research papers are an outdated format that slows the dissemination and effective use of knowledge. We advocate transitioning from plain-text narratives to richly annotated, graph-based representations in which nodes correspond to discrete logical units—such as hypotheses, experiments, methods, datasets, analyses, and conclusions —and the edges connect these nodes. Such structures will enhance readability for both humans and machines and, in the long term, help address several systemic problems in academic science, including the reproducibility crisis, the serials crisis, and weaknesses in peer review. In collaboration with our AI partner, we are developing a tool that converts existing text-based scientific documents into structured graph representations.
GeneBox: learning biology and having fun
As a team, we view mathematical modeling as a powerful tool for generating, validating, and refining scientific hypotheses. Within the broader field of evolutionary modeling, we are particularly inspired by models of gene regulatory networks and aim to use our modeling framework to facilitate understanding in microbiology and genetics. In collaboration with our educational partners, we are developing an interactive app that enables students to construct their own gene regulatory networks and explore resulting cell behaviors—learning core concepts through experimentation and play. We plan to integrate this tool into both school and university curricula.
Partners
Want more information? Follow Peter on Twitter X: @LidskyPeter
Our Team




Editing
Publications
Evolution of aging and diseases
- Lidsky^ PV, Yuan J, Andino^ R. Reconsidering life history theory amid infectious diseases. Trends in Ecology and Evolution. 2023. 38 (8):P699-700.
- Lidsky^ PV, Andino^ R. Could aging evolve as a pathogen control strategy? Trends in Ecology and Evolution. 2022. 37(12):1046-1057.
- Lidsky^ PV, Yuan J, Rulison JM, and Andino-Pavlovsky^ R. Is aging an inevitable characteristic of organic life or an evolutionary adaptation? Biochemistry (Moscow). 2022. 87(12): 1777–1817.
- Lidsky^ PV, Andino^ R. Epidemics as an adaptive driving force determining lifespan setpoints. Proc Natl Acad Sci U S A. 2020. 117(30):17937-17948.
- Lidsky^ PV, Andino R, Rouzine IM. Variability in viral pathogenesis: modeling the dynamic of acute and persistent infections. Curr Opin Virol. 2017. 23:120-124. PMID: 28551476. PMCID: PMC5695700
Virology
- mRNA-LNP vaccine-induced CD8+ T cells protect mice from lethal SARS-CoV-2 infection in the absence of specific antibodies. Montoya B, Melo-Silva CR, Tang L, Kafle S, Lidsky PV, Bajusz C, Vadovics M, Muramatsu H, Abraham E, Lipinszki Z, Chatterjee D, Scher G, Benitez J, Sung MMH, Tam YK, Catanzaro NJ, Schäfer A, Andino R, Baric RS, Martinez DR, Pardi N, Sigal LJ. Mol Ther. 2024 S1525-0016(24)00236-3.
- Escoubas CC, Dorman LC, Nguyen PT, Lagares-Linares C, Nakajo H, Anderson SR, Barron JJ, Wade SD, Cuevas B, Vainchtein ID, Silva NJ, Guajardo R, Xiao Y, Lidsky PV, Wang EY, Rivera BM, Taloma SE, Kim DK, Kaminskaya E, Nakao-Inoue H, Schwer B, Arnold TD, Molofsky AB, Condello C, Andino R, Nowakowski TJ, Molofsky AV. Type-I-interferon-responsive microglia shape cortical development and behavior. Cell. 2024 Apr 11;187(8):1936-1954.e24.
- Aviner R*, PV Lidsky*, Y Xiao, M Tasseto, L Zhang, PL McAlpine, J Elias, J Frydman, R Andino. SARS-CoV-2 Nsp1 regulates translation start site fidelity to promote infection. PLoS Pathogens. 2024. 9;20(2):e1011535.
- Yang* J, Xiao* Y, Lidsky* PV, Wu C-T, Bonser LR, Peng S, Garcia-Knight MA, Tassetto M, Chung C-I, Li X, Nakayama T, Lee IT, Nayak JV, Ghias K, Shoichet BK, Erle DJ, Jackson PK, Andino R, Shu X. Identifying natural compounds that potently inhibit SARS-CoV-2 in mouse models. Nat. Microbiology. 2023. 8:121–134.
- Wu* C-T, Lidsky* PV, Xiao* Y, Cheng* R, Lee IT, Nakayama T, Jiang S, He W, Demeter J, Garcia-Knight M, Turn RE, Rojas-Hernandez LS, Ye C, Chiem K, Shon J, Martinez-Sobrido L, Bertozzy CR, Nolan G, Milla C, Nayak JV, Andino R, Jackson PK. SARS-CoV-2 Replication in airway epithelia requires motile cilia and microvillar reprogramming. Cell. 2022. 186(1) 112-130.e20.
- Wu* C-T, Lidsky* PV, Xiao* Y, Lee* IT, Cheng* R, Nakayama* T, Jiang* S, Demeter J, Bevacqua RJ, Chang CA, Zhu B, Chen H, Goltsev Y, Tzankov A, Nayak JV, Nolan GP, Matter MS, Andino R, Jackson PK. SARS-CoV-2 infects human pancreatic beta-cells and elicits beta-cell impairment, Cell Metabolism. 2021. 33(8):1565-1576.e5.
- Xiao Y, Lidsky PV, Shirogane Y, Aviner R, Wu C-T, Li W, Zheng W, Talbot D, Catching A, Doitsh G, Su W, Gekko CE, Nayak A, Ernst JD, Brodsky L, Brodsky E, Rousseau E, Capponi S, Bianco S, Nakamura R, Jackson PK, Frydman J, Andino R. A defective viral genome strategy elicits broad protective immunity against respiratory viruses. Cell. 2021. 184(25):6037-6051.e14.
- Li* X, Lidsky* P, Xiao* Y, Wu C-T, Garcia-Knight M, Yang J, Nakayama T, Nayak JV, Jackson PK, Andino R, Shu X. Ethacridine inhibits SARS-CoV-2 by inactivating viral particles. PLoS Pathogens. 2021. 17(9): e1009898.
- Deng X, Garcia-Knight MA, Khalid MM, Servellita V, Wang C, Morris MK, Sotomayor-González A, Glasner DR, Reyes KR, Gliwa AS, Reddy NP, San Martin CS, Federman S, Cheng J, Balcerek J, Taylor J, Streithorst JA, Miller S, Sreekumar B, Chen P-Y, Schulze-Gahmen U, Taha TY, Hayashi J, Simoneau CR, Kumar GR, McMahon S, Lidsky PV, Xiao Y, Hemarajata P, Green NM, Espinosa A, Kath C, Haw M, Bell J, Hacker JK, Hanson C, Wadford DA, Anaya C, Ferguson D, Frankino, Haridha Shivram PA, Lareau LF, Wyman SK, Ott M, Andino R, Chiu CY. Transmission, infectivity, and neutralization of a spike L452R SARS-CoV-2 variant. Cell. 2021. 184(13):3426-3437.e8.
- Nakayama T; Lee I; Jiang S; Matter M; Yan C; Overdevest J; Goltsev Y; Shih L-C; Liao C-K; Wu C-T; Zhu B; Zarabanda D; Yang A; Schürch C; Chu P; Chen H; McIlwain D; Borchard N; Gall P; Dholakia S; Xu L; Stalder A; Lidsky PV; Xiao Y; Jackson P; Tai C-J; Yeh T-H; Andino R; Duran J; Mertz K; Patel Z; Haslbauer J; Menter T; Canoll P; DeConde A; Hwang P; Tzankov A; Nolan G; Nayak JV. Determinants of SARS-CoV2-2 entry and replication in airway mucosal tissue and susceptibility in smokers. Cell Reports Medicine. 2021. 2(10):100421.
- Krupina KA, Sheval EV, Lidsky^ PV. Variability in inhibition of host RNA synthesis by entero- and cardioviruses. J Gen Virol. 2010. 91(Pt 5):1239-44.
- Lidsky* PV, Romanova* LI, Kolesnikova MS, Bardina MV, Khitrina EV, Hato SV, van Kuppeveld FJ, Agol VI. Interactions between viral and prokaryotic pathogens in a mixed infection with cardiovirus and mycoplasma. J Virol. 2009. 83(19):9940-51. (Current topic in Microbe magazine, Nov 2009).
- Bardina* MV, Lidsky* PV, Sheval EV, Fominykh KV, van Kuppeveld FJ, Polyakov VY, Agol VI. Mengovirus-induced Rearrangement of the Nuclear Pore Complex: Hijacking Cellular Phosphorylation Machinery. J. Virol. 2009. 83(7):3150-61. (evaluated by F1000).
- Romanova* LI, Lidsky* PV, Kolesnikova MS, Fominykh KV, Gmyl AP, Sheval EV, Hato S, Van Kuppeveld FJM, Agol VI. Antiapoptotic Activity of the Cardiovirus Leader Protein, a Viral “Security” Protein. J. Virol. 2009. 83(14):7273-84.
- Lidsky* PV, Hato* S, Bardina* MV, Aminev AG, Palmenberg AC, Sheval EV, Polyakov VY, van Kuppeveld FJ, Agol VI. Nucleocytoplasmic traffic disorder induced by cardioviruses. J Virol. 2006. 80(6):2705-17. (spotlight in JVI, evaluated by F1000).
- Romanova LI, Belov GA, Lidsky PV, Tolskaya EA, Kolesnikova MS, Evstafieva AG, Vartapetian AB, Egger D, Bienz K, Agol VI. Variability in apoptotic response to poliovirus infection. Virology. 2005. 331(2):292-306.
- Belov* GA, Lidsky* PV, Mikitas OV, Egger D, Lukyanov KA, Bienz K, Agol VI. Bidirectional increase in permeability of nuclear envelope upon poliovirus infection and accompanying alterations of nuclear pores. J Virol. 2004. 78(18):10166-77.
Drosophila genetics and methods development
- Shekhar S, C Tracy, PV Lidsky, R Andino, KJ Wert, H Krämer. Sensory quiescence induces a cell-non-autonomous integrated stress response curbed by condensate formation of the ATF4 and XRP1 effectors. Nature communications. 2025. 16 (1), 252
- Lidsky PV^, J Yuan, KA Lashkevich, SE Dmitriev, R Andino^. Monitoring integrated stress response in live Drosophila. bioRxiv. 2023.07. 13.548942.
- Lidsky^ PV, Dmitriev SE, Andino^ R. Introduction of Dicistrovirus IRESes into UAS/SV40-polyA constructs results in premature polyadenylation and strong overexpression of the upstream ORF in Drosophila animals. bioRxiv. 2023.10.04.560905.
- Lidsky^ PV, Lukyanov KA, Misra T, Handke B, Mishin AS, Lehner CF. A genetically encoded fluorescent probe for imaging of oxygenation gradients in living Drosophila. Development. 2018. 145(4).
- Nayak A, Kim DY, Trnka MJ, Kerr CH, Lidsky PV, Stanley DJ, Rivera BM, Li KH, Burlingame AL, Jan E, Frydman J, Gross JD, Andino R. A Viral Protein Restricts Drosophila RNAi Immunity by Regulating Argonaute Activity and Stability. Cell Host Microbe. 2018. 24(4):542-557. (evaluated by F1000)
- Sarkisyan KS, Goryashchenko AS, Lidsky PV, Gorbachev DA, Bozhanova NG, Gorokhovatsky AY, Pereverzeva AR, Ryumina AP, Zherdeva VV, Savitsky AP, Solntsev KM, Bommarius AS, Sharonov GV, Lindquist JR, Drobizhev M, Hughes TE, Rebane A, Lukyanov KA, Mishin AS. Green fluorescent protein with anionic tryptophan-based chromophore and long fluorescence lifetime. Biophys J. 2015. 109(2):380-9.
- Handke B, Szabad J, Lidsky PV, Hafen E, Lehner CF. Towards long term cultivation of Drosophila wing imaginal discs in vitro. PLoS One. 2014. 9(9):e107333.
- Lidsky PV, Sprenger F, Lehner CF. Distinct modes of centromere protein dynamics during cell cycle progression in Drosophila S2R+ cells. J Cell Sci. 2013. 126 (20): 4782–4793.
* co-first author, ^ corresponding or co-corresponding author
26 January 2026