- 1A-207, 2/F, Block 1, To Yuen Building
- +852 3442-2199
- +852 3442-0549
- zongli.zheng@cityu.edu.hk
- CityUHK Scholars
- Gene Rearrangement • Liquid Biopsy • Precision Genome Editing
Prof. Zheng received his PhD degree with distinction in medical epidemiology from Karolinska Institutet in 2011. He then completed his postdoctoral training at Massachusetts General Hospital and Harvard Medical School, supported by the Swedish Research Council International Postdoc Fellowship. In July 2015, he joined the Department of Biomedical Sciences at City University of Hong Kong.
Research Interests & Contribution
We are interested in genomic medicine and biotechnology innovation to advance our understanding, diagnosis and treatment of diseases. We invented the AMP technology, which has significantly accelerated gene fusion discovery. Initially implemented in top U.S. hospitals, the AMP technology has since been adopted globally in molecular diagnosis assays to guide targeted therapy for cancer patients. We co-developed the GUIDE-seq method, a widely used reference for genome-wide unbiased profiling of CRISPR edits. GUIDE-seq has supported the first CRISPR therapy approved by FDA in December 2023 that utilizes the precise genome targeting technology CRISPR/Cas9.
Recently, we revealed gene rearrangement or fusion heterogeneity-associated therapy resistance in non-small cell lung cancer patients, suggesting that tailored/precision targeting approaches are needed for treating these cancer patients. We observed in patient tumors, under the ground, numerous “futile” rearrangements involving genes that are in close proximity within a cell nucleus, leading us to introduce the concept “passenger fusion”. To address the “passenger fusion” issue before targeted therapy, we recommend using orthogonal diagnosis methods beyond DNA-based approaches. Subsequently, we developed an ultra-sensitive clinical-grade RNA-seq-based rearrangement calling algorithm SplitFusion. On genome editing, we developed GenomePAM, a novel tool enabling direct identification of PAM sequences in mammalian cells. This approach achieves greater than 10,000-fold higher efficiency while being simpler than previous in-cell methods. Additionally, we have discovered new CRISPR/Cas nucleases from nature and, leveraging AI models, engineered variants with desired PAM preferences for therapeutics.
Currently, our research employs multi-disciplinary approaches, including genomics, AI-powered protein engineering, and molecular epidemiology, and centring on:
- Lung Cancer: to understand the molecular heterogeneity that underpins resistances to targeted therapy and immunotherapy in lung cancer patients.
- Precision Genome Editing: to advance precision genome editing and delivery technologies for the development of safe and curative treatments.
- Liquid Biopsy: to identify molecular determinants and circulating signatures that can be used for cancer monitoring, diagnosis, and early detection.
Funding
We are grateful for the financial supports from City University of Hong Kong, Ming Wai Lau Centre of Reparative Medicine / Karolinska Institutet, Innovation and Technology Commission of Hong Kong, Swedish Research Council, National Natural Science Foundation of China, Research Grant Council of Hong Kong, and Shenzhen Medical Research Fund.
Position Available
We are looking for motivated individuals who are passionate about genomic medicine to join our lab as research assistants, PhD students and postdocs.
Innovation and Collaborations
-
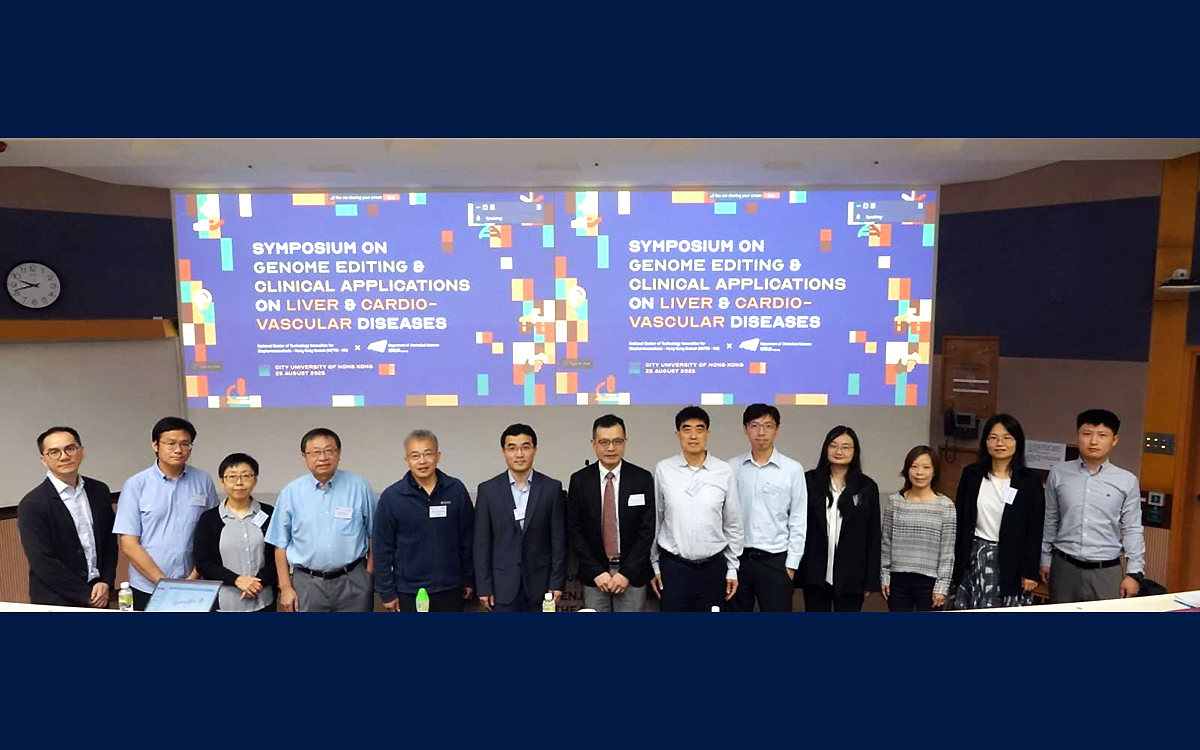 Symposium on Genome Editing & Clinical Applications on Liver & Cardiovascular Diseases
Symposium on Genome Editing & Clinical Applications on Liver & Cardiovascular Diseases -
 RAISe+ project: In Vivo Somatic Human Genome Editing for Genetic Diseases: Translating Novel Genome Editing and Engineered Delivery Vectors into Clinical Trials
RAISe+ project: In Vivo Somatic Human Genome Editing for Genetic Diseases: Translating Novel Genome Editing and Engineered Delivery Vectors into Clinical Trials -
 InFocus: Cell & Gene Therapy in Hong Kong
InFocus: Cell & Gene Therapy in Hong Kong -
 Genome Editing experts from Beijing and Shanghai (北京大學,清華大學,復旦大學,華東師範大學), oversea (Harvard University, King Saud bin Abdulaziz University, Ajou University School of Medicine, Karolinska Institutet), and local universities (香港大學,香港中文大學,香港科技大學,香港城市大學) unite in Hong Kong
Genome Editing experts from Beijing and Shanghai (北京大學,清華大學,復旦大學,華東師範大學), oversea (Harvard University, King Saud bin Abdulaziz University, Ajou University School of Medicine, Karolinska Institutet), and local universities (香港大學,香港中文大學,香港科技大學,香港城市大學) unite in Hong Kong
Latest News





Publications
(Google Scholar full list: 65 publications cited > 17,000 times as of 1 January 2025)
Five selected recent publications (since 2021)
- Yu M, Ai L, Wang B, Lian S, Ip L, Liu J, Li L, Tsai SQ, Kleinstiver BP, Zheng Z#. GenomePAM directs PAM characterization and engineering of CRISPR-Cas nucleases using mammalian genome repeats. Nature Biomedical Engineering (2025) [Link]
(Shortly after GenomePAM appeared online, it was illustrated by an incredible AI-Powered Drug Discovery Platform [8-min video] despite not being 100% accurate.)
- Bian W*, Zhang B*, Song Z*, Knisbacher BA*, Chan YM, Bao Y, Xu C, Wang W, Chu HY, Lu C, Wang H, Bao S, Gong Z, Keung HY, Chow ZY, Zhang Y, Cheuk W, Getz G, Nardi V, Yang M, Cho WCS#, Wang J#, Chen J#, Zheng Z#. SplitFusion enables ultrasensitive gene fusion detection and reveals fusion variant-associated tumor heterogeneity. Patterns (2025) [Link]
- Lian S, Lu C, Li F, Yu X, Ai L, Wu BH, Gong X, Zhou W, Liang X, Zhan J, Yuan Y, Fang F, Liu Z#, Ji M#, Zheng Z#. Monitoring hepatocellular carcinoma using tumor content in circulating cell-free DNA. Clinical Cancer Research (2024) 30: 2772–2779. [Link]
- Song Z*, Lian S*, Mak S*, Chow ZY*, Xu C, Wang W, Keung HY, Lu C, Kebede TF, Gao Y, Cheuk W, Cho CS, Yang M, Zheng Z#. Deep RNA sequencing revealed fusion junctional heterogeneity may predict crizotinib treatment efficacy in ALK-rearranged non-small cell lung cancer. Journal of Thoracic Oncology (2022) 17:264-276. [Link]
- Song Z, Lu C, Xu C, Zheng Z#. Noncanonical gene fusions detected at the DNA level necessitate orthogonal diagnosis methods before targeted therapy [Invited commentary]. Journal of Thoracic Oncology (2021) 16:344-348. [Link]
Five selected postdoctoral studies (Advisor: Dr John Iafrate; Collaboration lab: Dr Keith Joung)
- Kleinstiver BP, Pattanayak V, Prew MS, Tsai SQ, Nguyen NT, Zheng Z, Joung JK. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature (2016) 529:490–495.
- Tsai SQ*, Zheng Z*, Nguyen NT, Liebers M, Topkar VV, Thapar V, Wyvekens N, Khayter C, Iafrate AJ, Le LP, Aryee MJ, Joung JK. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nature Biotechnology (2015) 33:187–198.
- Kleinstiver BP, Prew MS, Tsai SQ, Nguyen NT, Topkar VV, Zheng Z, Joung JK. Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nature Biotechnology (2015) 33:1293-1298.
- Shaw A, Ou S, Bang Y, Camidge R, Solomon B, Salgia R, Riely G, Varella-Garcia M, Shapiro G, Costa D, Doebele R, Le LP, Zheng Z, Tan W, Stephenson P, Shreeve S, Tye L, Christensen J, Wilner K, Clark J, Iafrate AJ. Crizotinib in ROS1-Rearranged Non-Small Cell Lung Cancer. The New England Journal of Medicine (2014) 371:1963-1971.
- Zheng Z, Liebers M, Zhelyazkova B, Cao Y, Panditi D, Lynch KD, Chen J, Robinson HE, Shim HS, Chmielecki J, Pao W, Engelman JA, Iafrate AJ, Le LP. Anchored multiplex PCR for targeted next-generation sequencing. Nature Medicine (2014) 20:1479-1484.
Five selected doctoral studies (Advisor: Dr Weimin Ye)
- Zheng Z#, Advani A, Melefors O, Glavas S, Nordstrom H, Ye W, Engstrand L, Andersson AF. Titration-free 454 sequencing using Y adapters. Nature Protocols (2011) 6:1367-1376.
- Zheng Z#, Advani A, Melefors O, Glavas S, Nordstrom H, Ye W, Engstrand L, Andersson AF. Titration-free massively parallel pyrosequencing using trace amounts of starting material. Nucleic Acids Research (2010) 38:e137.
- Zheng Z, Jia Y, Hou L, Persson C, Yeager M, Lissowska J, Chanock S, Blaser M, Chow WH, Ye W. Genetic Variation in A4GNT in Relation to Helicobacter pylori Serology and Gastric Cancer Risk. Helicobacter (2009) 14:120–125.
- Zheng Z, Margolis KL, Liu S, Tinker LF, Ye W, Women's Health Initiative I. Effects of estrogen with and without progestin and obesity on symptomatic gastroesophageal reflux. Gastroenterology (2008) 135:72-81.
- Zheng Z, Nordenstedt H, Pedersen NL, Lagergren J, Ye W. Lifestyle factors and risk for symptomatic gastroesophageal reflux in monozygotic twins. Gastroenterology (2007) 132:87-95.
31 May 2026

![Shortly after GenomePAM appeared online, it was illustrated by an incredible AI-Powered Drug Discovery Platform [8-min video] despite not being 100% accurate.](/bms/img/profile/zlz/media-20250813-genomepam.jpg)