MoRNiNG




About the MoRNiNG database
The MoRNiNG The MoRNiNG (Modification of RNAs in Natural rG4) database provides information on RNA modifications located within experimentally confirmed RNA G-quadruplex (rG4) structures or putative G-quadruplex-forming sequences (PQSs). These sequences are derived from mRNA, lncRNA, and miRNA (both pre-miRNA and mature-miRNA) across nine species: human, mouse, rhesus monkey, chimpanzee, pig, rat, zebrafish, HIV-1, and SV40. The database includes details on nine RNA modifications: m6A, m1A, m7G, m5C, m6Am, m5U, A-to-I, 2’-O-Me, and Ψ.
The MoRNiNG The MoRNiNG (Modification of RNAs in Natural rG4) database provides information on RNA modifications located within experimentally confirmed RNA G-quadruplex (rG4) structures or putative G-quadruplex-forming sequences (PQSs). These sequences are derived from mRNA, lncRNA, and miRNA (both pre-miRNA and mature-miRNA) across nine species: human, mouse, rhesus monkey, chimpanzee, pig, rat, zebrafish, HIV-1, and SV40. The database includes details on nine RNA modifications: m6A, m1A, m7G, m5C, m6Am, m5U, A-to-I, 2’-O-Me, and Ψ.
RNA G-quadruplex
RNA G-quadruplexes (rG4s) are non-canonical secondary structures formed by the stacking of two or more G-quartets— planar arrangements held together by Hoogsteen hydrogen bonds among four guanines. Specifically, putative G-quadruplex-forming sequences (PQSs) typically contain four G-tracts separated by three loops, a pattern that enables computational prediction. RNA modifications occurring within or near these PQSs have the potential to influence rG4 thermostability, interactions with rG4-binding proteins, and conformational dynamics. Given that many PQSs are nested within canonical stem-loop structures in the transcriptome, exploring the role of RNA modifications in regulating conformational dynamics is essential for understanding RNA function.
RNA G-quadruplexes (rG4s) are non-canonical secondary structures formed by the stacking of two or more G-quartets— planar arrangements held together by Hoogsteen hydrogen bonds among four guanines. Specifically, putative G-quadruplex-forming sequences (PQSs) typically contain four G-tracts separated by three loops, a pattern that enables computational prediction. RNA modifications occurring within or near these PQSs have the potential to influence rG4 thermostability, interactions with rG4-binding proteins, and conformational dynamics. Given that many PQSs are nested within canonical stem-loop structures in the transcriptome, exploring the role of RNA modifications in regulating conformational dynamics is essential for understanding RNA function.
Framework
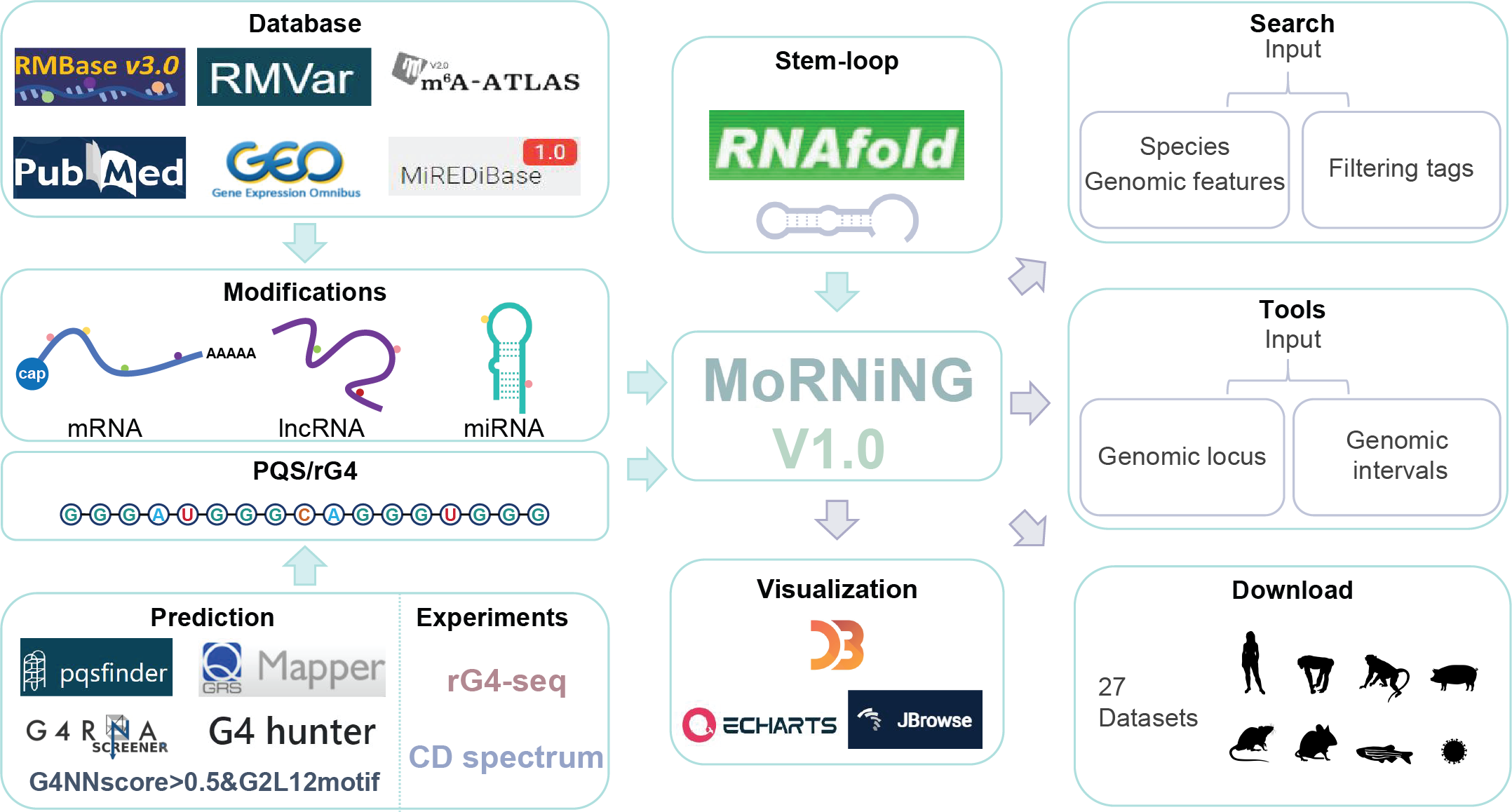

Data collection & processing
The MoRNING database integrates RNA modification sites from six sources, including GEO, RMbase, RMVar, MiREDiBase, m6A Atlas, and published datasets. Here we used seven software to identify PQSs in the longest transcripts, which include both mRNA and lncRNA, based on transcript biotypes from the Ensembl database (release 109) and miRNAs from miRBase (version 22). The software used includes pqsfinder (score > 1), QGRS (default parameter: score >= 17), G4hunter (-w 25 -s 1.2), G4RNA screener (cGcC >= 4.5, G4H >= 0.9, G4NN >= 0.5), RNAfold, and G4NN score > 0.5 & Quadparser-based (G >= 2N1-12)3G >= 2 sequence motif (G2L12 motif). Additionally, the database also features PQSs with experimental support from techniques including rG4-seq, ultra-low-input rG4-seq, and circular dichroism spectrum.
For each entry in the database, potential stem-loop structures are predicted using RNAfold, with the results displayed in dot-bracket notation. However, rG4/PQS lacking stem-loop structures will not show the dot-bracket illustration.
Furthermore, we collected 801 potential GI-Quadruplex-forming sequences (PIQS) on human transcripts for rG4 introduced by A-I editing, which can be obtained using the "ModType" option on the search page.
The MoRNING database integrates RNA modification sites from six sources, including GEO, RMbase, RMVar, MiREDiBase, m6A Atlas, and published datasets. Here we used seven software to identify PQSs in the longest transcripts, which include both mRNA and lncRNA, based on transcript biotypes from the Ensembl database (release 109) and miRNAs from miRBase (version 22). The software used includes pqsfinder (score > 1), QGRS (default parameter: score >= 17), G4hunter (-w 25 -s 1.2), G4RNA screener (cGcC >= 4.5, G4H >= 0.9, G4NN >= 0.5), RNAfold, and G4NN score > 0.5 & Quadparser-based (G >= 2N1-12)3G >= 2 sequence motif (G2L12 motif). Additionally, the database also features PQSs with experimental support from techniques including rG4-seq, ultra-low-input rG4-seq, and circular dichroism spectrum.
For each entry in the database, potential stem-loop structures are predicted using RNAfold, with the results displayed in dot-bracket notation. However, rG4/PQS lacking stem-loop structures will not show the dot-bracket illustration.
Furthermore, we collected 801 potential GI-Quadruplex-forming sequences (PIQS) on human transcripts for rG4 introduced by A-I editing, which can be obtained using the "ModType" option on the search page.
Functions
The MoRNING database offers three major functional modules: search, tool, and download. Users can search for rG4 structures featuring various RNA modification sites across different species, customized to their specific needs. Alternatively. Users can manually input genomic loci or intervals, and utilize our tool function to directly find predicted PQS on the tool page. For convenience, we offer 27 datasets from nine different species for users to download.
The MoRNING database offers three major functional modules: search, tool, and download. Users can search for rG4 structures featuring various RNA modification sites across different species, customized to their specific needs. Alternatively. Users can manually input genomic loci or intervals, and utilize our tool function to directly find predicted PQS on the tool page. For convenience, we offer 27 datasets from nine different species for users to download.
Manuals
Search-mainpage:
This function enables users to efficiently query PQS with modifications within a specific genome range for a given species.
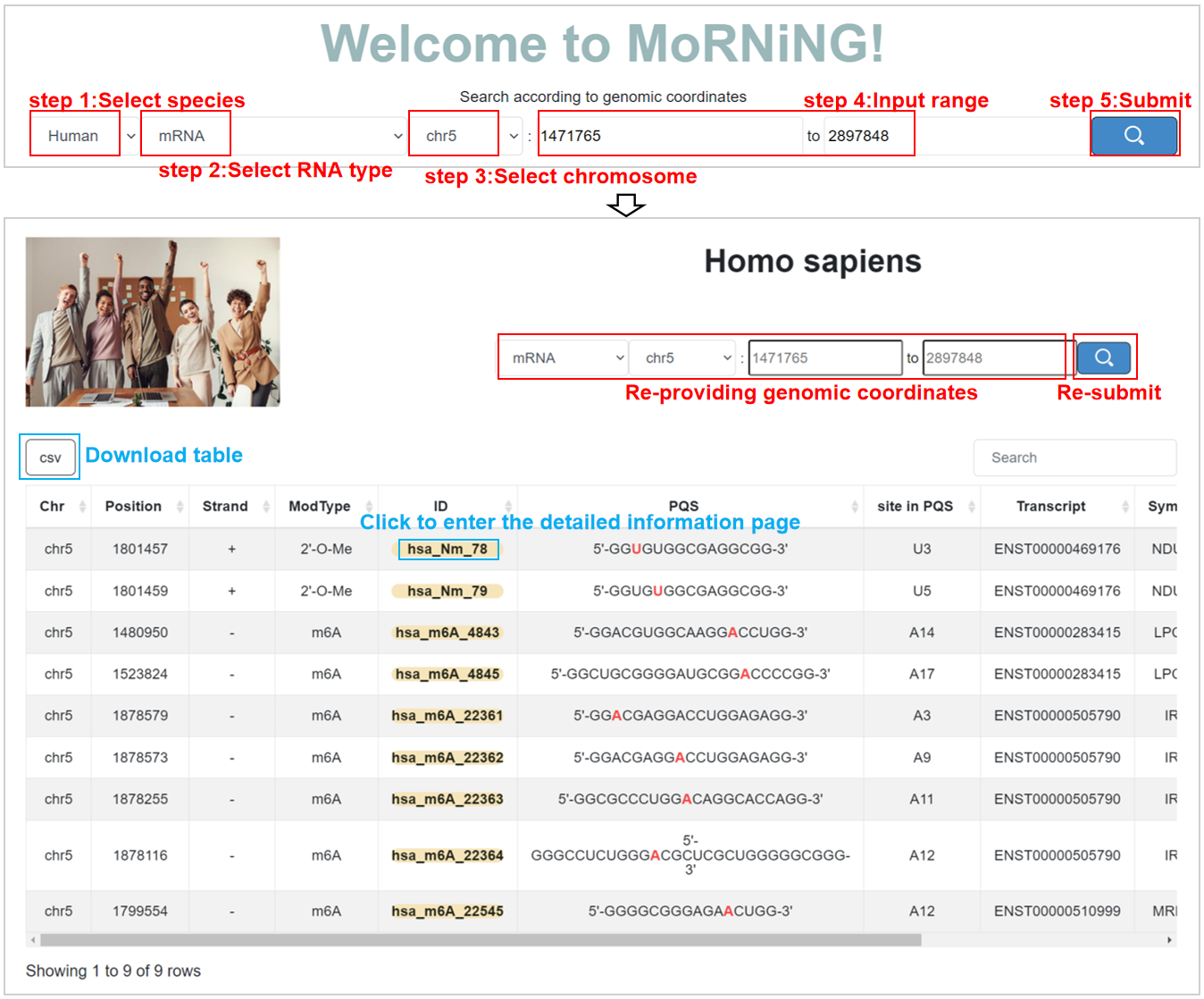
Search:
Users can use multiple filter tags to query PQS with modifications. The above four options are required.
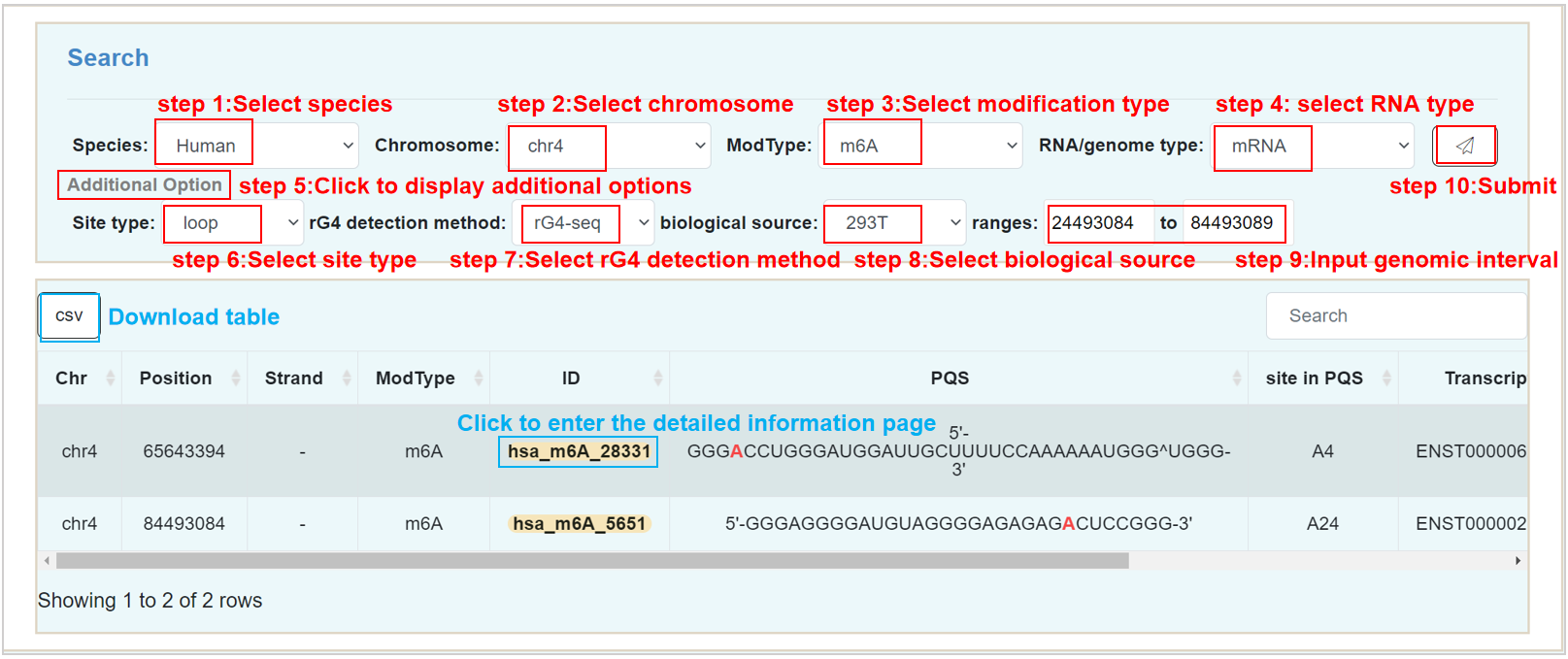
Genomic loci:
Users can provide the locus on the exon of a species to find all the intersected PQS.
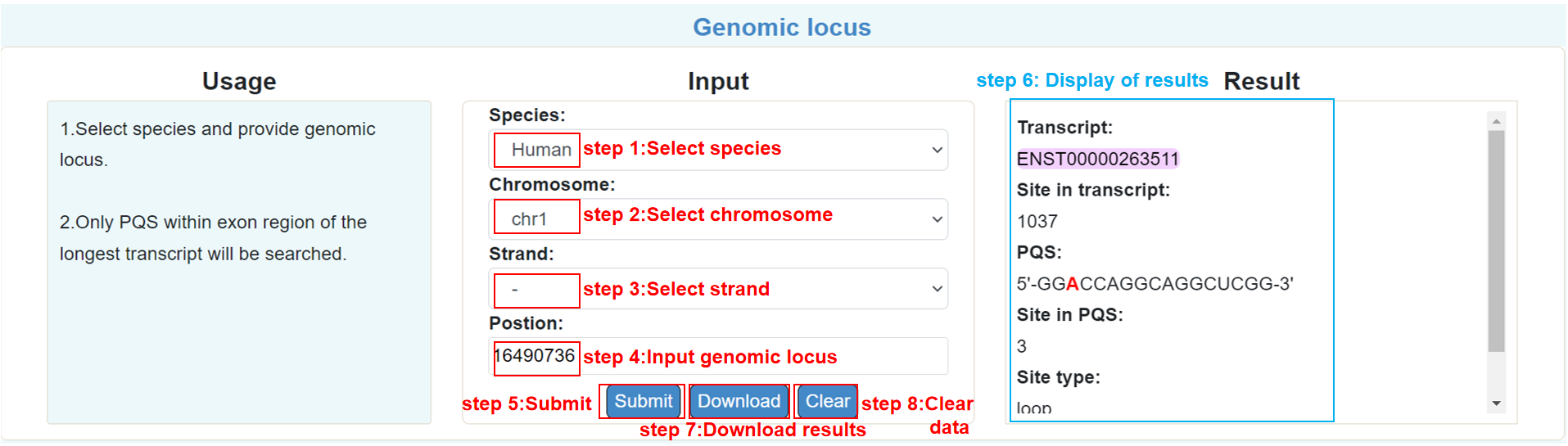
Genomic interval:
By providing the genomic intervals, users can identify the predicted PQSs.
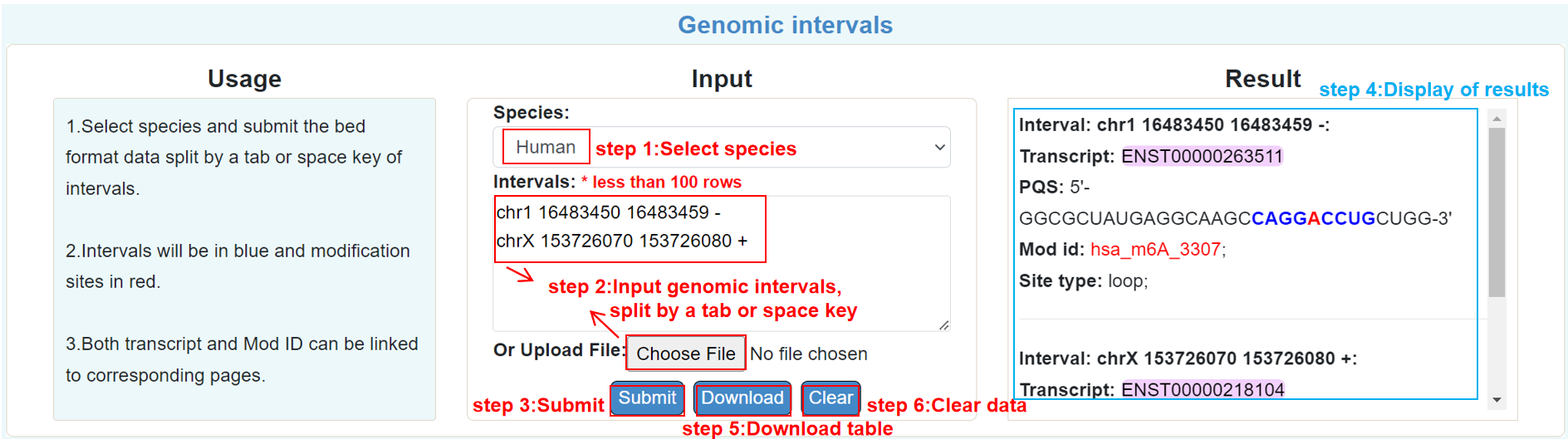
Search-mainpage:
This function enables users to efficiently query PQS with modifications within a specific genome range for a given species.

Search:
Users can use multiple filter tags to query PQS with modifications. The above four options are required.

Genomic loci:
Users can provide the locus on the exon of a species to find all the intersected PQS.

Genomic interval:
By providing the genomic intervals, users can identify the predicted PQSs.

Detection of Modification
| Category | Method | literature | DOI |
| High | Aza-IP | Nat Biotechnol | 10.1038/nbt.2566 |
| RNA-BisSeq | Nucleic Acids Res | 10.1093/nar/gkn954 | |
| BS-seq-with-improved-protocol | Nat Struct Mol Biol | 10.1038/s41594-019-0218-x | |
| DART-seq | Nat Methods | 10.1038/s41592-019-0570-0 | |
| DART-seq&Bullseye | Mol Cell | 10.1016/j.molcel.2021.12.038 | |
| CeU-Seq | Nat Chem Biol | 10.1038/nchembio.1836 | |
| Nm-REP-seq | Sci China Life Sci | 10.1007/s11427-022-2210-0 | |
| CIMS_miCLIP | Nat Methods | 10.1038/nmeth.3453 | |
| CITS_miCLIP | Nat Methods | 10.1038/nmeth.3453 | |
| m6ACE-seq | Nat Commun | 10.1038/s41467-019-13561-z | |
| miCLIP | Nat Methods | 10.1038/nmeth.3453 | |
| m6A-REF-seq | Sci Adv | 10.1126/sciadv.aax0250 | |
| m6A-SAC-seq | Nat Protoc | 10.1038/s41596-022-00765-9 | |
| m6A-seq-with-improved-protocol | Cell Rep | 10.1016/j.celrep.2014.05.048 | |
| DM-Ψ-seq | Nat Chem Biol | 10.1038/s41589-019-0420-5 | |
| FICC-Seq | Nucleic Acids Res | 10.1093/nar/gkz658 | |
| m6A-CLIP/IP | Genes Dev | 10.1101/gad.269415.115 | |
| PA-m6A-Seq | Methods Mol Biol | 10.1007/978-1-0716-1374-0_8 | |
| m6A-Label-seq | STAR Protoc | 10.1016/j.xpro.2021.101096 | |
| GLORI | Nat Biotechnol | 10.1038/s41587-022-01487-9 | |
| PRAISE | Nat Chem Biol | 10.1038/s41589-023-01304-7 | |
| BACS | Nat Methods | 10.1038/s41592-024-02439-8 | |
| m1A-IP-seq | Nat Methods | 10.1038/s41592-019-0550-4 | |
| m1A-quant-seq | Nat Methods | 10.1038/s41592-019-0550-4 | |
| m1A-MAP | Mol Cell | 10.1016/j.molcel.2017.10.019 | |
| UBS-seq | Nat Biotechnol | 10.1038/s41587-023-02034-w | |
| Nm-Mut-seq | Cell Res | 10.1038/s41422-023-00836-w | |
| Nm-seq | Nat Methods | 10.1038/nmeth.4294 | |
| RBS-Seq | Proc Natl Acad Sci U S A | 10.1073/pnas.1817334116 | |
| RiboMeth-seq | Methods Mol Biol | 10.1007/978-1-4939-6807-7_13 | |
| m7G-miCLIP-Seq | Cell Res | 10.1038/s41422-019-0230-z | |
| m7G-seq | Mol Cell | 10.1016/j.molcel.2019.03.036 | |
| MAZTER-seq | Cell | 10.1016/j.cell.2019.06.013 | |
| meCLIP | RNA | 10.1261/rna.078543.120 | |
| miCLIP&m6Aboost | Nucleic Acids Res | 10.1093/nar/gkab485 | |
| miCLIP2&m6Aboost | Nucleic Acids Res | 10.1093/nar/gkab485 | |
| RNA-Seq | Nat Methods | 10.1038/s41592-018-0017-z | |
| Ψ-seq | Cell | 10.1016/j.cell.2014.08.028 | |
| Slic-seq | Nucleic Acids Res | 10.1093/nar/gkad604 | |
| Medium | m1A-seq | Nature | 10.1038/nature16998 |
| m5C-RIP-seq | Methods Mol Biol | 10.1007/978-1-4939-9045-0_24 | |
| m6A-Seal-seq | Nat Chem Biol | 10.1038/s41589-020-0525-x | |
| m6A-seq | Nature | 10.1038/nature11112 | |
| miR-mmPCR-seq | Genome Res | 10.1101/gr.224386.117 | |
| BoRed-seq&m7G-RIP-seq | Mol Cell | 10.1016/j.molcel.2019.03.040 | |
| m6A-CLIP | Nature | 10.1038/nature21355 | |
| Low | DARNED | Bioinformatics | 10.1093/bioinformatics/btq285 |
| RADAR | Nucleic Acids Res | 10.1093/nar/gkt996 | |
| Modomics | Nucleic Acids Res | 10.1093/nar/gkab1083 | |
| ELIGOS | Nucleic Acids Res | 10.1093/nar/gkaa620 | |
| Prediction by CNN model | Nucleic Acids Res | 10.1093/nar/gkaa811 | |
| Protocol for analyzing RNA editing in mature miRNAs using deep sequencing data | Methods Mol Biol | 10.1007/978-1-4939-2291-8_14 |
Detection of rG4
| Category | Method | literature | DOI |
| Experiment | rG4-seq | Nat Methods | 10.1038/nmeth.3965 |
| Ultra-low-input rG4-seq | Under Review at Nature Portfolio | 10.21203/rs.3.rs-5687952/v1 | |
| circular dichroism spectrum | NA | NA | |
| Prediction | pqsfinder | Bioinformatics | 10.1093/bioinformatics/btx413 |
| QGRS | Hum Genomics | 10.1186/1479-7364-8-8 | |
| G4Hunter | Bioinformatics | 10.1093/bioinformatics/btz087 | |
| G4RNA screener | Biochimie | 10.1016/j.biochi.2018.06.002 | |
| G4NNscore>0.5&G2L12motif | Chem Commun (Camb) | 10.1039/c8cc04635b |
Citation
Zhou Y, Lyu S, Liew SW, Mou X, Hoffecker I, Yan J, Li Y, Kwok CK, Zhang J. MoRNiNG: A Database of RNA Modification Sites Associated with RNA Secondary Structure Dynamics. Genomics Proteomics Bioinformatics. 2025 Nov 14:qzaf106.
Zhou Y, Lyu S, Liew SW, Mou X, Hoffecker I, Yan J, Li Y, Kwok CK, Zhang J. MoRNiNG: A Database of RNA Modification Sites Associated with RNA Secondary Structure Dynamics. Genomics Proteomics Bioinformatics. 2025 Nov 14:qzaf106.